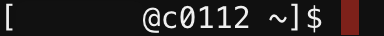Getting Started¶
Cheaha is the batch compute cluster platform of the Research Computing System (RCS). Cheaha is intended primarily for batch processing of research computing software. We offer a user-friendly portal website Open OnDemand with graphical interfaces to the most common features, all in one place. Read on to learn more about our resources and how to access them.
Cheaha is appropriate for the following types of workflows:
- High Performance Computing (HPC)
- High Throughput Computing (HTC)
- Batch Computing
- Graphical User Interface (GUI) and other interactive software
Please bear in mind the following expectations when using Cheaha.
- Resources are shared by many researchers.
- Jobs may take time to start.
- Jobs, partitions, and researchers all have caps on total resources in use at any one time.
- All jobs have a maximum time limit.
If your needs go beyond these expectations, please Contact Support.
What if I Need Help?¶
Please Contact Us with requests for support. Tips on getting effective support are available in the Support Guide, and answers to common questions can be found in our section for Frequently Asked Questions (FAQ).
Account Creation¶
Please visit our Account Creation page for detailed instructions on creating a Cheaha account.
Accessing Cheaha¶
The primary method for accessing Cheaha is through our online portal website, Open OnDemand. To login to our portal, navigate to our https://rc.uab.edu, which does not require an on-campus connection nor the UAB Campus VPN. You should be presented with UAB's Single Sign-on page, which will require use of Duo 2FA. Login using the appropriate credentials laid out at our Account Creation page.
SSH may be used to access Cheaha. Connect to host cheaha.rc.uab.edu on port 22.
Enabling SSH Key Authentication on Cheaha¶
SSH key authentication allows you to log in to Cheaha from your personal computer (PC) using the command-line interface (CLI) without entering a password each time. This improves security and enables passwordless login.
SSH Key Setup on Cheaha¶
If you have already set up your SSH keys on your local machine, see Local Machine CLI Setup, you can enable passwordless login on Cheaha as follows:
-
Add your public key to authorized_keys:
-
First, log in to Cheaha. You can do this either: Using a terminal with SSH and your password from you PC (as shown below) Or via OOD Cheaha shell access.
-
After you login in to Cheaha, append your ECDSA public key to authorized_keys:
This ensures that your public key is registered on the server so future SSH logins can use passwordless authentication.
-
-
Set correct permissions
After adding your public key to authorized_keys, you need to make sure both the authorized_keys file and the .ssh directory have the correct permissions. This is important because SSH will refuse to use files or directories that are accessible by other users, which can prevent passwordless login from working.
-
Test SSH login After adding your public key and setting the correct permissions, you should verify that SSH key authentication works correctly. This ensures that future logins to Cheaha, including OOD Cheaha shell access will not prompt for a password.
With Integrated Development Environments (IDEs)¶
An alternative method suited for developers is to use IDEs like VSCode, using the "Remote - Tunnels" extension, to connect to an HPC Desktop Interactive Job. You can find more information for setting this up in the VSCode Tunnel section.
Please do not use IDEs like VSCode "Remote - SSH" or Cursor Server to connect to Cheaha. These methods run all of their processes on the login node, which are automatically terminated. Instead, use "Remote - Tunnel" as described in the VSCode Tunnel section. Please read additional information about working on the login node. See examples of processes that are automatically terminated on the login node, please note they are not limited to the items in the table below.
| Technology | Process | Acceptable | Result |
|---|---|---|---|
| VS Code with Remote - Tunnels" | Runs from a compute node | ✔️ Yes | Process runs normally |
| VS Code with Remote - SSH" | Runs on a login node | ❌ No | Process is terminated |
| Cursor Server | Runs on a login node | ❌ No | Process is terminated |
Open OnDemand Features¶
The Open OnDemand portal features a terminal usable directly in the browser for basic functions such as file management.
Interactive apps are available directly in your browser, including the following.
More detailed documentation can be found on our Open OnDemand page.
Hardware¶
A full list of the available hardware can be found on our hardware page.
Storage¶
All researchers are granted 5 TB of individual storage when they create their Research Computing account.
Shared storage is available to all Lab Groups and Core Facilities on campus. Shared storage is also available to UAB Administration groups.
Please visit our Storage page for detailed information about our individual and shared storage options.
Partitions¶
Compute nodes are divided into groups called partitions each with specific qualities suitable for different kinds of workflows or software. In order to submit a compute job, a partition must be chosen in the Slurm options. The partitions can be roughly grouped as such:
| Use | Partition Names | Notes |
|---|---|---|
| GPU Processing | pascalnodes, pascalnodes-medium, amperenodes, amperenodes-medium | These are the only partitions with GPUs |
| All Purpose | amd-hdr100 | Runs AMD CPUs compared to all other CPU partitions running Intel. Contact us with issues running on this partition |
| Shorter time | express, short, intel-dcb | |
| Medium-long time | medium, long | |
| Very large memory | largemem, largemem-long |
Please visit our hardware for more details about the partitions.
Etiquette¶
Quality-of-Service (QoS) limits are in place to ensure any one user can't monopolize all resources.
Why You Should Avoid Running Jobs on Login Nodes¶
To effectively manage and provide high-performance computing (HPC) resources to the University community provided via the clusters, kindly use the terminal from compute nodes in created jobs rather than the terminal from login nodes. Our clusters are essential for conducting this large and complex scientific computations that often times require a significant amount of computing power. These clusters are shared environments, where multiple users execute their research and computing tasks simultaneously. It is important to utilize the structure of these environments properly for efficient and respectful use of the shared resources, so everyone gets a fair chance at using these resources.
Login vs. Compute Nodes¶
Like with most HPC clusters, cheaha nodes are divided into two, the login node and compute nodes. The login node acts as the gateway for users to access the cluster, submit jobs, and manage files. Compute nodes, on the other hand, are like the engines of the cluster, designed to perform the heavy lifting of data processing and computation.
The Login node can be accessed from the Cheaha landing page or through the $HOME directory. You can see in the images below, how to identify if you’re within a login node or compute node.
You are on the login node if:
- terminal prompt looks like
[$USER@login004 ~]$
You are on compute nodes if:
- using Open OnDemand Interactive Apps
- using Interactive Jobs with
srun - terminal prompt looks like
[$USER@c0112 ~]$
Important
If the terminal prompt appears as bash-4.2$ instead of the user prompt [$USER@login004], please refer to the FAQ below to resolve the issue.
How to Restore Default Terminal Prompt From bash-4.2$ to $USER@login004?¶
There might be scenarios where you see the terminal prompt display bash-4.2$ like below, instead of the user prompt [$USER@login004].
The bash-4.2$ prompt indicates that the files $HOME/.bashrc and/or $HOME/.bash_profile are missing or corrupted. To resolve this issue, we recommend following these steps:
(i) If you have made any changes to these files earlier, it's advisable to create backups. For instance, you can rename .bashrc to .bashrc.backup, and you can verify if the backed-up files are listed.
bash-4.2$ mv $HOME/.bashrc $HOME/.bashrc.backup
bash-4.2$ mv $HOME/.bash_profile $HOME/.bash_profile.backup
bash-4.2$ ls .bash*
.bash_profile.backup .bashrc.backup
(ii) After you have taken the backup of those files, run the following command to copy the default versions from /etc/skel to $HOME. Doing this will clobber, or remove, any changes you may have made, so be sure to make a backup first, as shown in the step (ii), if you wish to keep any changes. You will see the copied files listed in the directory.
bash-4.2$ cp /etc/skel/.bash* $HOME
bash-4.2$ ls .bash*
.bash_profile .bash_profile.backup .bashrc .bashrc.backup
(iii) You can exit the terminal or source your files using the commands below to apply the changes, after which you will see the user prompt.
Slurm and Slurm Jobs¶
Slurm Workload Manager is a widely used open-source job scheduler that manages the queue of jobs submitted to the compute nodes. It ensures efficient use of the cluster by allocating resources to jobs, prioritizing tasks, and managing queues of pending jobs. Starting Slurm jobs can be done in two primary ways: using Open OnDemand (OOD) or through the terminal. For more details on how to use Slurm on cheaha, please see our slurm docs.
What Should Run in Jobs?¶
Ideally, only non-intensive tasks like editing files, or managing job submissions should be performed on the login node. Compute-intensive tasks, large data analyses, and simulations should be submitted as Slurm jobs to compute nodes. This approach ensures that the login node remains responsive and available for all users to manage their tasks and submissions. Submitting compute-intensive tasks as Slurm jobs to compute nodes helps to prevent overloading the login node, ensuring a smoother experience for all users of the cluster.
How to Start Slurm Jobs?¶
There are two straightforward ways to start Slurm jobs on cheaha, and they are detailed below.
Open OnDemand (OOD)¶
UAB uses the OOD platform, a web-based interface for providing access to cluster resources without the need for command-line tools. Users can easily submit jobs, manage files, and even use interactive applications directly from their browsers. One of the standout features of OOD is the ability to launch interactive applications, such as a virtual desktop environment. This feature allows users to work within the cluster as if they were on a local desktop, providing a user-friendly interface for managing tasks and running applications. For an overview of how the page works, and to read more details see our docs on Navigating Open OnDemand. After logging into OOD, users can access various applications designed for job management, file editing, and more.
Terminal (sbatch Jobs)¶
For users comfortable with the command line, submitting jobs via scripts using sbatch is a straightforward process. An sbatch script contains the job specifications, such as the number of nodes, execution time, and the command to run. This method provides flexibility and control over job submission and management. For more information on this, please see our docs on Submitting Jobs with Slurm.
Important
If you are doing more than minor file management, you will need to use a compute node. Please request an interactive session at https://rc.uab.edu or through a job submitted using Slurm.
Slurm¶
Slurm is our job queueing software used for submitting any number of job scripts to run on the cluster. We have documentation on how to set up job scripts and submit them further in. More complete documentation is available at https://slurm.schedmd.com/.
Software¶
A large variety of software is available on Cheaha as modules. To view and use these modules see the following documentation.
For new software installation, please try searching Anaconda for packages first. If you still need help, please send a support ticket
Conda Packages¶
A significant amount of open-source software is distributed as Anaconda or Python libraries. These libraries can be installed by the user without permission from Research Computing using Anaconda environments. To read more about using Anaconda virtual environments see our Anaconda page.
If the software installation instructions tell you to use either conda install or pip install commands, the software and its dependencies can be installed using a virtual environment.
Frequently Asked Questions¶
-
What should I do to access shared storages and recognize my group membership after being added to a group on Cheaha?
-
Do you have any processes/connections on
cheaha.rc.uab.edu?- Please exit and log back in.
- If you have active Tmux/Screen sessions, you will need to terminate those as well, log out, log back in and start Tmux.
-
Do you have an active Open OnDemand session?
- In Open OnDemand (https://rc.uab.edu), navigate to the green navigation bar in the top right corner. Look for the
HelporDeveloperdropdown menu and click on it. Then, clickRestart Web Server. Once the restart is complete, please try again.
- In Open OnDemand (https://rc.uab.edu), navigate to the green navigation bar in the top right corner. Look for the
-
Do you have one or more OOD HPC Desktops running?
- Terminate the desktops and start new ones.
-
-
Why should I use SSH key–based login for Cheaha?
SSH key–based login not only allows passwordless access but also helps avoid account lockouts or cool-off periods caused by multiple incorrect password attempts. It provides a more secure and reliable way to connect to Cheaha. For setup instructions, see Enabling SSH Key Authentication on Cheaha.
How to Get Help¶
For questions, you can reach out via our various channels.